icon96™
Stop gambling on cycle numbers. The icon96 uses iconPCR™ technology with AutoNorm™ software to provide real-time fluorescence feedback in each of 96 independent wells, preventing under-amplification dropouts and over-amplification artifacts—delivering balanced, sequencing-ready libraries without manual quantification or normalization.
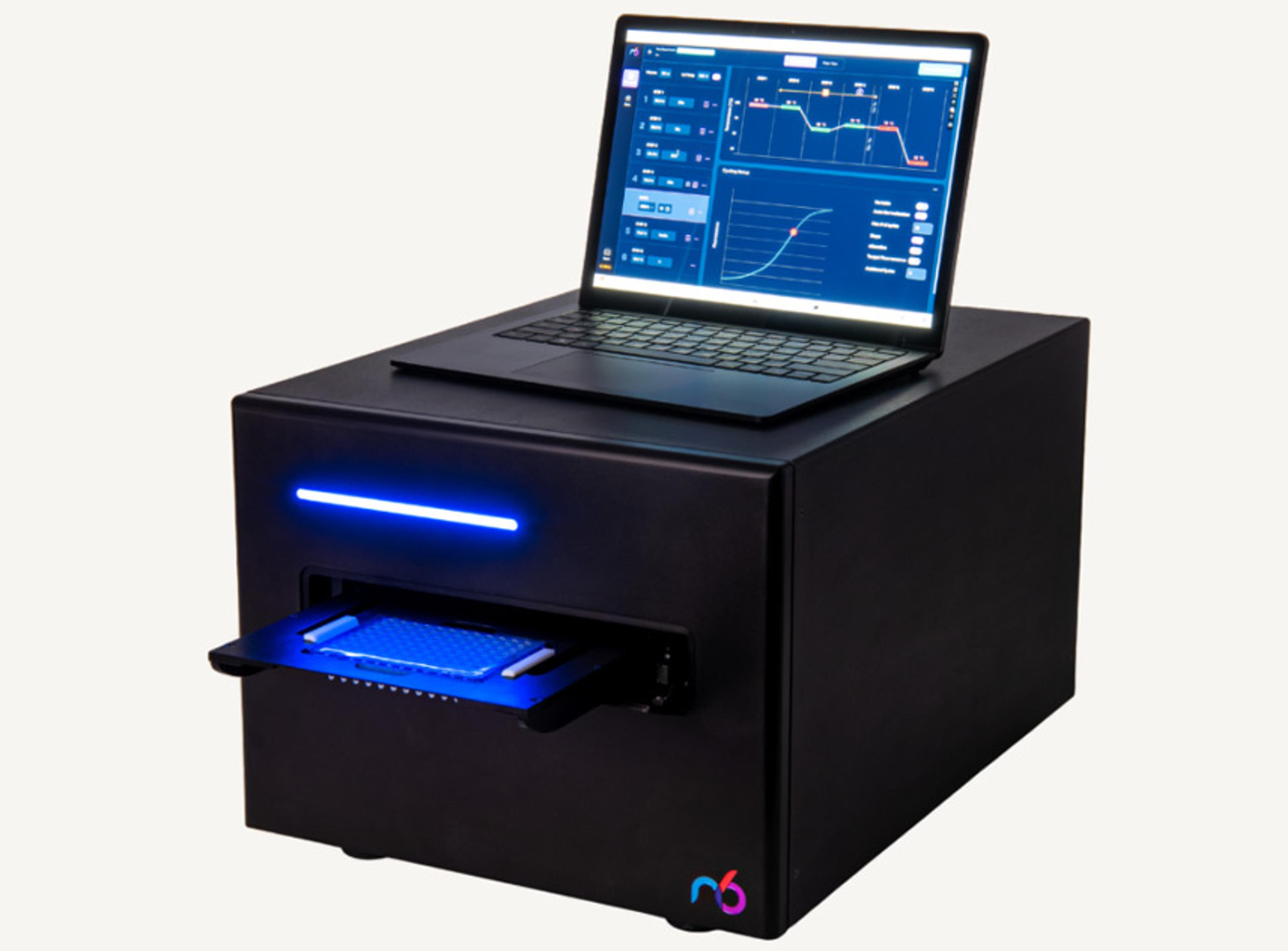
Every NGS library prep risks under-amplifying precious samples into dropouts or over-amplifying them into chimera-filled garbage data—and you won't know until it's too late. The icon96 solves this with iconPCR™ technology: 96 individually controlled thermal zones with real-time fluorescence detection. AutoNorm™ software monitors every well and stops each reaction at its optimal cycle, ensuring data quality and workflow efficiency in one automated step.
Improve Sequencing Data Quality:
- Eliminate dropouts: Real-time monitoring prevents under-amplification failures—critical for precious clinical samples like FFPE, cfDNA, or biopsies you can't re-collect
- Reduce artifacts: AutoNorm stops reactions before over-cycling creates PCR duplicates, chimeras, or adapter dimers that compromise sequencing data quality
- Rescue variable-quality samples: Automatically adjusts cycles for degraded or low-input samples (DIN 1.3–8), avoiding both failed libraries and over-amplification distortions
Relieve Library Prep Workflow Bottlenecks:
- 96 → 1 cleanup: Pool libraries by volume directly after PCR; perform one SPRI cleanup instead of 96 (saves ~$225/plate)
- No manual normalization: AutoNorm delivers every sample at the same target fluorescence—sequence-ready without Qubit quantification, TapeStation analysis, or dilution steps
- Flexible throughput: Run RNA-seq, 16S metagenomics, single-cell, WGS, or FFPE workflows (and more coming) on the same instrument with any existing kit—no proprietary consumables required
- Time savings: Complete library prep to sequencing pool in under 3 hours versus 8+ hours with traditional workflows
How It Works:
The icon96 features 96 miniaturized thermocyclers with precision temperature control and real-time fluorescence detection in every well (iconPCR technology). AutoNorm monitors library amplification every cycle and automatically stops individual wells when they reach optimal yield—preventing both under- and over-amplification across variable inputs (0.6 ng to 1000 ng) without prior sample quantification.
The Bottom Line - Why it's iconic:
- 96 wells, each with independent control
- AutoNorm: No more manual quant or normalization
- Every sample gets its perfect cycling—no batching
- Real-time feedback, automatic cycle adjustment
- Save low-input and tough samples with fewer failures
- One simple cleanup—ditch individual SPRIs
- Works with your kits and workflows—no “locked-in” extras
- Faster prep, lower costs, better data quality
Brochures
Scale your NGS operations with icon96
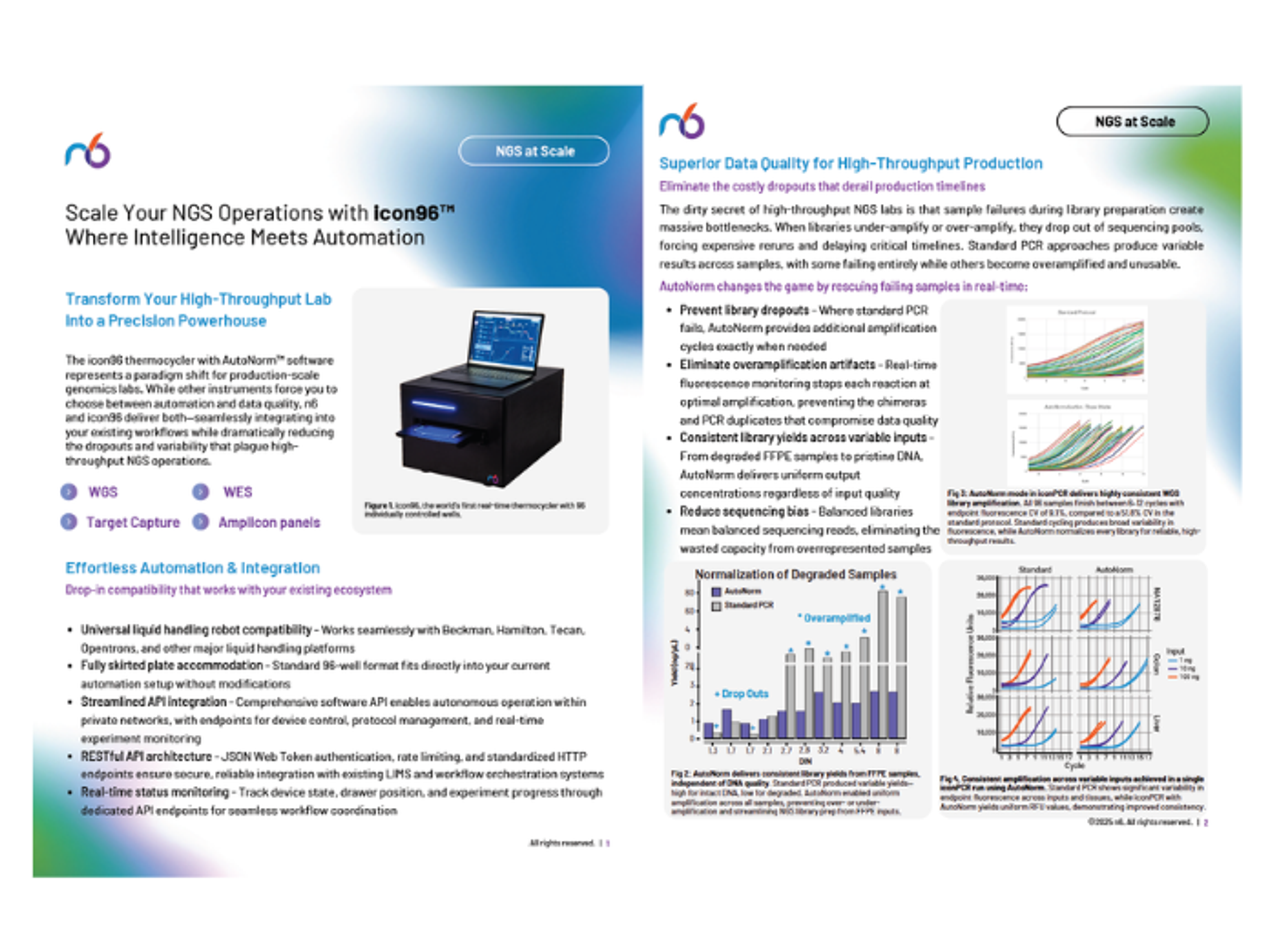
Find out how to transform your high-throughput lab into a precision powerhouse with the icon96 thermocycler with AutoNorm™ software from n6. This solution promises both automation and data quality, seamlessly integrating into your existing workflows while reducing the dropouts and variability that plague high-throughput NGS operations.
icon96 - Amplify, quantify, and normalize all at once
Learn more about iconPCR™ from n6, the world’s first thermocycler with 96 individually controlled wells and built-in AutoNorm™ software, designed for normalized, sequencing-ready libraries in one step.
Streamlined FFPE DNA library prep using NEBNext UltraShear and Multiplex Oligos kits with iconPCR
n6 Tec and NEB offer a streamlined FFPE DNA library preparation workflow that enhances amplification consistency, reduces hands-on time by up to 60%, and maintains high data quality across variable inputs.
This application note highlights an optimized workflow using NEBNext UltraShear and NEBNext Multiplex Oligos with iconPCR AutoNorm, which dynamically tunes PCR cycling, minimizes batching and QC burden, and delivers uniform, high-quality NGS libraries from challenging FFPE samples.
How to optimize low-input cfDNA library prep for targeted sequencing sensitivity
Wednesday, March 25 at 15:00 GMT | 16:00 CET | 11:00 EDT | 8:00 PDT
Register now to get practical NGS workflow strategies to boost sensitivity, consistency, and confidence!
Cell-free DNA (cfDNA) sequencing can enable highly sensitive detection of low allele-fraction variants, but achieving reliable performance at low input requires careful control of library preparation and hybrid capture. PCR amplification is a key source of variability that can affect yield, coverage uniformity, and ultimately assay sensitivity and reproducibility.
In this SelectScience® webinar, held in partnership with n6 Tec, we will discuss practical strategies for optimizing cfDNA workflows and demonstrate how icon96 supports assay development and production by enabling sample-specific PCR conditions.
Attendees will learn how tailored amplification can improve consistency across precious samples, support balanced pooling, and strengthen confidence in downstream variant detection.
Key learning objectives:
- Identify key technical challenges in targeted sequencing from low-input cfDNA.
- Explain how PCR amplification during library preparation impacts sensitivity, accuracy, and reproducibility.
- Evaluate how sample-specific PCR optimization using iconPCR96 can improve workflow consistency in development and production.
Who should attend?
This webinar is ideal for genomics professionals, core lab managers, and translational researchers working with cfDNA sequencing for MRD detection, liquid biopsy workflows, or low-input targeted panels who need to optimize PCR amplification for improved assay performance and reproducibility.
Certificate of attendance
If you attend the live webinar, you will automatically receive a certificate of attendance, including a learning outcomes summary, for continuing education purposes.
If you view the on-demand webinar, you can request a certificate of attendance by emailing editor@selectscience.net.
Webinar details
- Cost: Free to attend
- Location: Online
- Duration: 60 minutes
Registration is required to secure your place. If you register but can’t attend live, you will receive a link to the on‑demand recording once it becomes available.